
POWERED BY THE TRUTH LOOP
THE SAINT ECOSYSTEM
Modular Intellegence for the modern laboratory
The SAINT Ecosystem is a modular AI platform built for laboratory and research environments. It integrates with digital imaging systems and laboratory information workflows, processes data through specialized analysis modules, and presents results through structured interfaces designed for review.
Vendor-agnostic and configurable, SAINT is designed to operate within existing infrastructure and support analytical workflows across laboratory and research settings.
DUAL ENGINE SHOWCASE
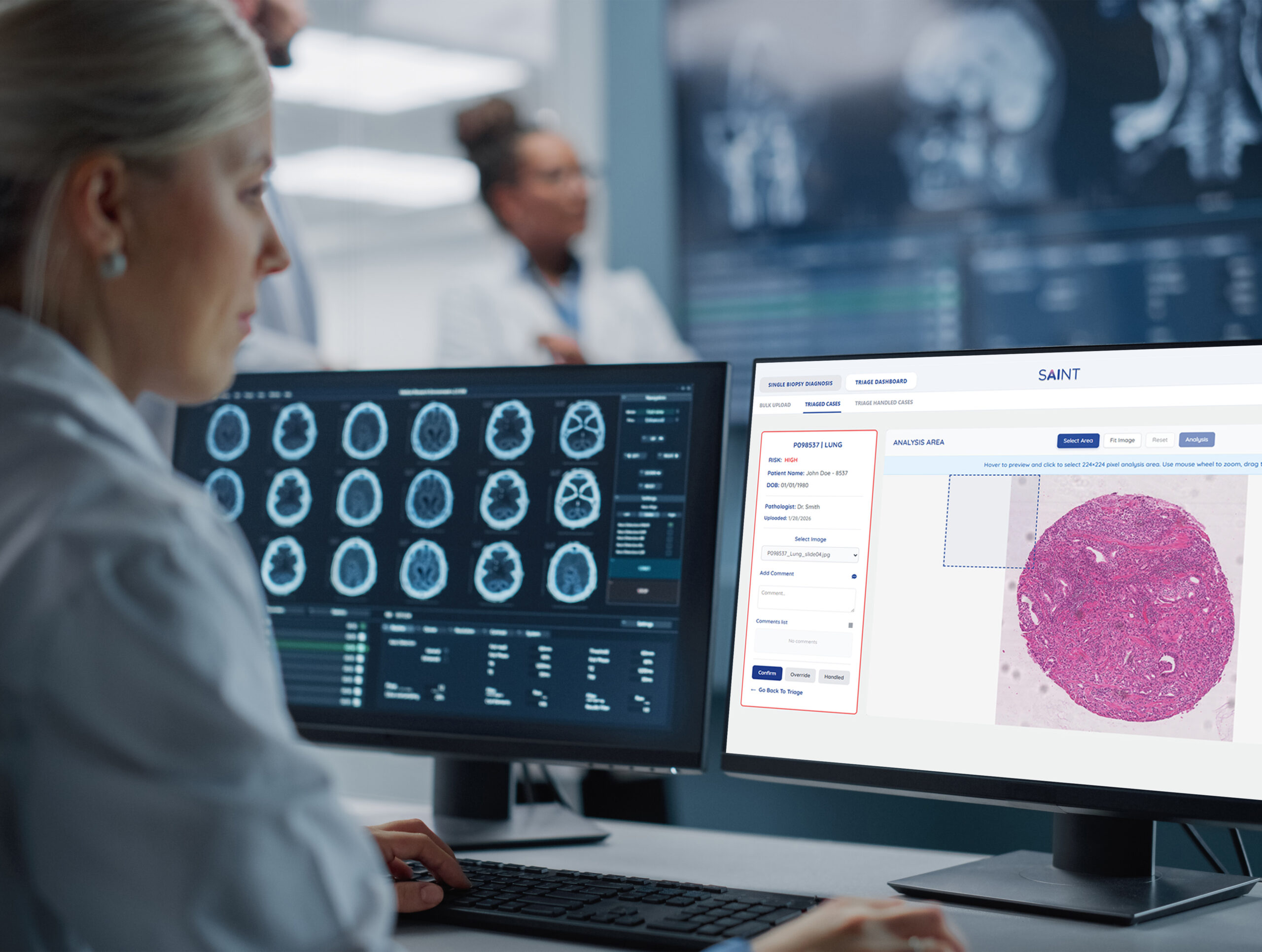
The Pathology Suite
Automated Workflow Support & Case Prioritization
Internal testing demonstrated workflow time reduction in controlled settings.
- Designed to highlight cases requiring additional review
Smart-Snip ROI: Pathologist-defined analysis. Draw a box around any region of interest to instantly run AI quantification on that specific area.
The Discovery Suite
Biomarker Analysis & Cohort Exploration
- Quantitative Histology: No black boxes. Measure TILs and Budding precisely.
- Support exploratory identification of immune-pattern subgroups for further study.
- Longitudinal Validation: Trained on 5-year outcomes, not just morphology
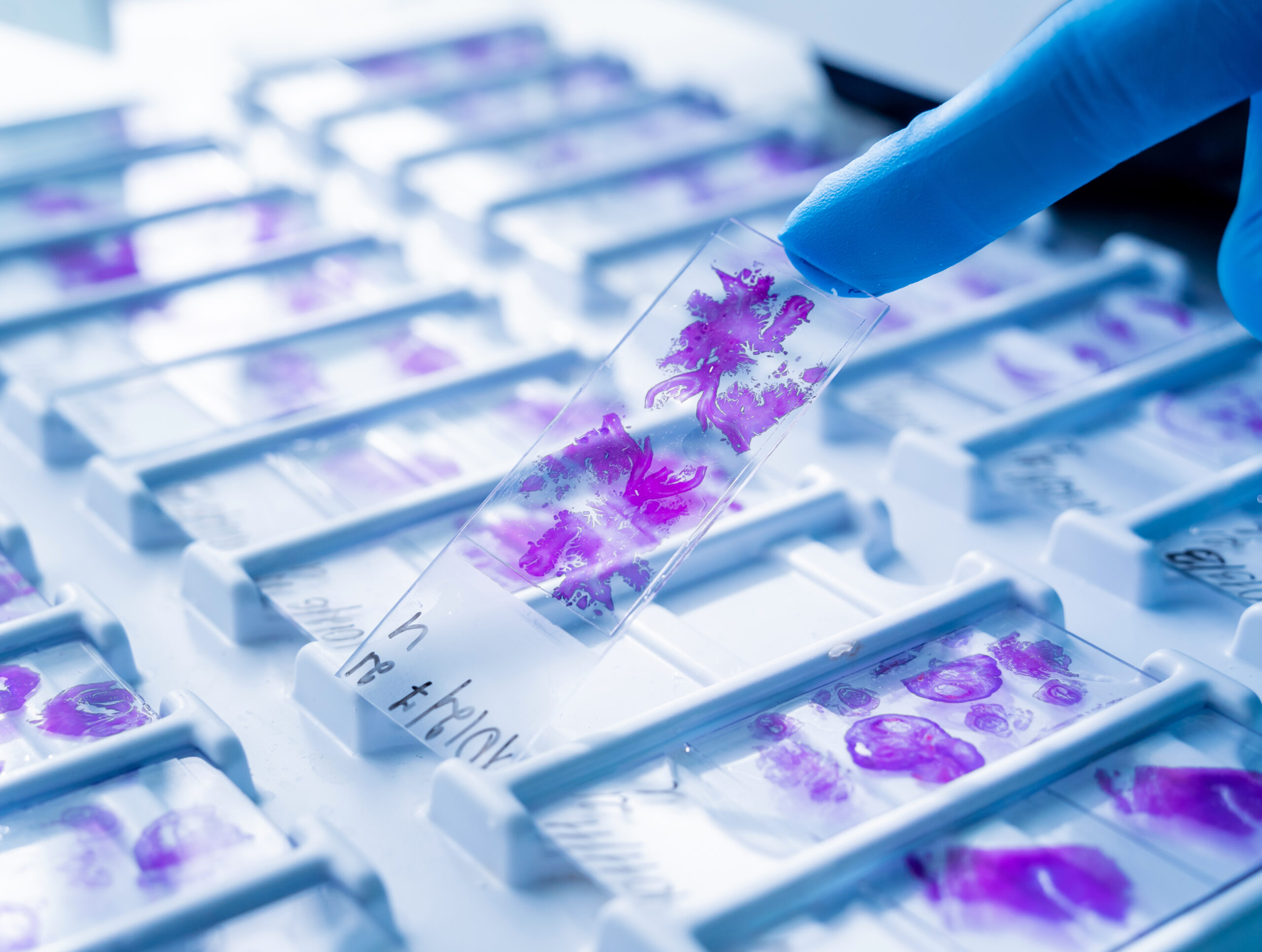

THE DEPLOYMENT SPECS
Vendor-Agnostic Middleware
Designed to integrate with common WSI scanners and LIS environments using a middleware layer, without requiring full system replacement.
High-Performance Rendering
Optimized architecture supports low-latency visual overlays within standard digital pathology viewing environments.
Security & Data Controls
Built with security and privacy controls designed for healthcare environments, including configurable data handling policies.
Shadow Mode Monitoring
Parallel Model Evaluation
When configured, the system can operate in parallel evaluation mode, comparing model outputs with pathologist review and associated data inputs for internal performance analysis.
THE DEPLOYMENT MATRIX
For Clinical Laboratories

Deploy the Pathology Suite
- Pilot Program: 30-day evaluation period using your existing digital pathology workflow.
- Supports integration with common interoperability standards (e.g., HL7, FHIR), depending on deployment configuration.
- Supports integration with common interoperability standards (e.g., HL7, FHIR), depending on deployment configuration.
For Pharmaceutical Partners

Access the Discovery Suite
- Retrospective analysis using available longitudinal datasets.
- Support exploratory cohort identification based on configured biomarker criteria.
- Support development of proprietary research assays and analytical tools.
PRODUCT & TECHNICAL FAQ.
Clinical & Regulatory
Technical & Workflow

The Longitudinal Data Platform.
Connecting clinical pathology data and research workflows to support oncology discovery and translational research.
Email: info@santoviapath.ai
Location: Boston, MA
Copyright © 2026 Santovia PathAI. All rights reserved. | Privacy Policy | Terms of Service